Plasmid
Plasmids are extrachromosomal DNA molecules. They are small, circular and have the ability to replicate autonomously. Replication of plasmid is not under the control of chromosomal DNA. They are mostly found in bacteria. Some of the eukaryotes like yeast and plants also contain plasmids.
Download Complete Chapter Notes of Biotechnology: Principles and Processes
Download Now
Their ability to replicate independently makes plasmid a cloning vector in the recombinant DNA technology for transferring and manipulating genes.
- Many antibiotic-resistant genes in bacteria are present in plasmids.
- The size of plasmid varies from a few base pairs to thousands of bp.
- Plasmids also get transferred from one bacterial cell to another by the process of conjugation.
- Plasmids carrying a specific gene are introduced into bacterial cells, which multiply rapidly and the required DNA fragment is produced in larger quantities.
- Plasmids are used to prepare recombinant DNA with the desired gene to transfer genes from one organism to another. This is known as genetic engineering.
- Joshua Lederberg coined the term plasmid.
Table of Content
Recommended Video:

Plasmid Structure
- Plasmids are extrachromosomal and not essential. They are useful but not necessarily present in every organism of the species
- Plasmids are not a part of the genome and the same plasmid can exist in different species and gets transferred from one another
- Plasmids have their own origin of replication (ORI) and they replicate along with the cell so that each daughter cell possesses a copy of the plasmid also
- Apart from the origin of replication, often it contains genes for antibiotic resistance, for the production of toxins and other useful genes, that may be required for the survival of cells
| Don’t miss: Answer Key NEET 2022 |
Plasmid Vector
Plasmids and bacteriophages are frequently used as cloning vectors in DNA recombinant technology.
- The ease with which plasmids can be modified and replicated makes it a great tool in genetic engineering and biotechnology
- For genetic engineering purposes, plasmids are artificially prepared in the lab
- The lab-grown plasmids, which are used as a vector contain an origin of replication, cloning site and selection marker
| Vector Element | Description |
| Origin of Replication
(ORI) |
DNA sequence where initiation of replication starts |
| Selectable Marker | For selecting bacteria containing desired plasmid, e.g. antibiotic resistance genes and other specific genes |
| Multiple Cloning Sites (MCS) | Recognition sites to insert foreign DNA fragment by using restriction enzymes, a few or single recognition site is preferred to avoid getting several fragments |
| Promoter Region | Promotes transcription of the target gene to get the desired protein |
| Primer Binding site | The sequence of DNA used as a start point for PCR amplification and sequence verification |
- DNA is cut at the specific points by using restriction enzymes (molecular scissors), which make sticky ends of the DNA
Herbert Boyer and Stanley Norman Cohen together discovered recombinant DNA technology by recombining DNA segments as desired and inserting them into the bacteria cell to get the desired protein
- The desired genes are then inserted by using DNA Ligase
- The recombinant DNA molecule is then introduced to the host bacteria cell by the process of transformation
- The recombinant plasmid then multiplies using host DNA polymerase
- The first plasmid used as a cloning vector was pSC101 of Salmonella typhimurium. They showed that a gene from a frog can be expressed in the bacterial cell
- E.coli. plasmid is frequently used as a cloning vector
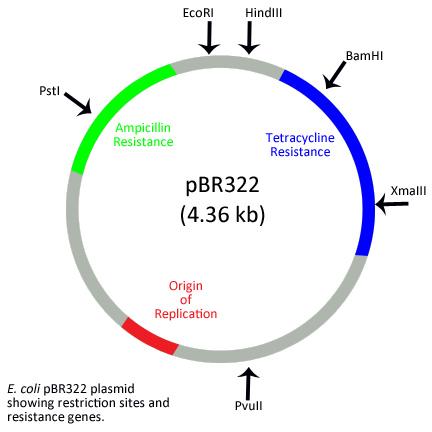
pBR322 Plasmid
The main characteristics of pBR322 are:
- Restriction sites: BamH I, Hind III, Sal I, Pvu I, Pvu II, Pst I, EcoR I, Cla I
- Selectable marker: antibiotic resistance genes for ampicillin (ampR) and tetracycline (tetR)
- ORI: the origin of replication
- ROP: It codes for proteins, which are involved in the process of replication of plasmid
Different antibiotic resistance genes act as a restriction site and to ligate foreign DNA and for the selection of transformants. The gene, where the foreign DNA is inserted becomes inactive.
Alternative selectable marker: Mostly these have the ability to produce some colour after reacting with a chromogenic substance. The alternative markers are used for the ease of differentiating recombinants from non-recombinants, e.g. gene coding for β-galactosidase.
When a foreign gene is inserted between the gene coding for β-galactosidase, the recombinant cell does not produce the enzyme β-galactosidase due to inactivation of the gene. In the presence of a chromogenic substrate, non-recombinants form blue colour colonies and recombinants form colourless colonies.
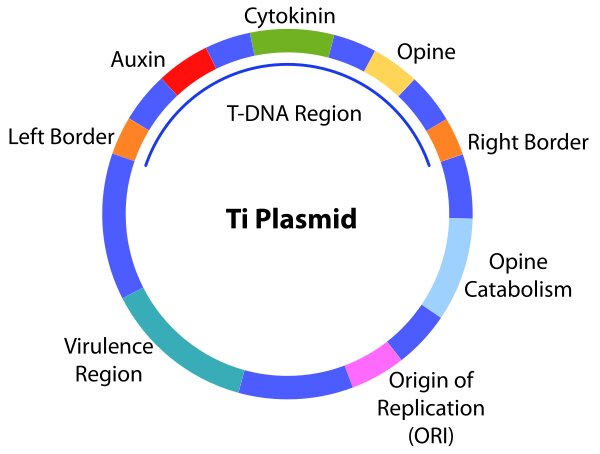
Ti Plasmid
The Tumour inducing or Ti plasmid is present in the bacterium Agrobacterium tumifaciens.
It is widely used now as a cloning vector to deliver desirable genes to the host plant to get transgenic plants. The main characteristics of Ti plasmid are:
- Size of the plasmid is ~ 250kbp
- There are different kinds of Ti plasmids based on the different genes they possess, which code for different opines, e.g. leucinopine, nopaline, octopine, etc.
- It is a pathogenic species to many dicotyledonous plants. It causes crown gall disease in plants.
- It contains one or more T-DNA region
- Agrobacterium tumifaciens has the ability to transform the normal cells into tumour cells by inserting a DNA piece known as T DNA and it starts producing chemicals, that are required by the bacterium
- After inserting the desired gene into Ti plasmid, it loses its pathogenic ability but is still able to insert the desired gene into the plant cell
- It contains vir or virulence genes, which transfer T-DNA region to plant cells and gets integrated into the plant genome
- Ti plasmid can be modified as per the requirement to insert the desired genes
- Agrobacterium tumifaciens is called “nature’s genetic engineer”
Recombinant Plasmid
Recombinant plasmids are the plasmids into which a foreign DNA fragment of gene is inserted. These recombinant plasmids independently replicate from the chromosomal DNA of the host.
Cells that contain recombinant plasmids usually are selected by taking to advantage the antibiotic resistance marker found on the vector plasmid. The cells containing recombinant plasmids can be recognized as containing recombinant plasmids by screening for insertional inactivation of a second genetic marker on the plasmid.
Advantages of Plasmids
- Plasmids are used as tools to transfer, manipulate and clone genes
- Since bacteria can rapidly divide, it can be used as factories to copy DNA fragments in large quantities
- Using plasmids as vectors is advantageous as it is easy to isolate and manipulate because of small size
- Due to its circular configuration, it is more stable
- Replicate independant of the host
Learn more about various concepts in biotechnology related to NEET only at BYJU’S.
Frequently Asked Questions
What is an extrachromosomal DNA?
Any DNA that is present outside the chromosome is called an extrachromosomal DNA. It can be either outside or inside a cell nucleus. Examples – Mitochondrial DNA, Plasmids.
Plasmid has been used as vector because?
Plasmids can be used as tools to transfer, clone and manipulate genes. Those plasmids that are used experimentally for such purposes are called vectors. Into plasmid vectors, DNA fragments or genes can be inserted to create a “recombinant plasmid”. Thus, they are used as vectors as it can carry foreign DNA fragments when it is introduced into them.
Who discovered plasmid?
Name the plasmid present in Agrobacterium tumefaciens.
What is a bacteriophage?
It is a virus that infects bacteria and it also replicates within the bacterial cell. It is composed of proteins that encapsulate an RNA or DNA genome. It is also referred to as a phage.
What does pBR322 stand for?
The letter p stands for plasmid. BR denotes the name of two researchers, Bolivar and Rodriguez. The number 322 denotes the order of synthesis that distinguishes it from other plasmids synthesized in the same laboratory.
Further reading:

Comments